Biotechnology of probiotics and intestinal pathogens
The intestine plays a crucial role in our body, not only in absorbing nutrients from the food we eat but also in hosting a complex network of interactions involving intestinal enterocytes (cells lining the intestine), microbiome bacteria (microorganisms living in the gut), and the immune system. Changes in these factors can disrupt the stability of these interactions, allowing harmful bacteria to invade the intestine and cause diseases.
In our research laboratory, we focus on studying how pathogenic bacteria utilize a specialized mechanism called the Type 3 Secretion System to infect the intestine and evade the immune system's defenses. These bacteria employ specific proteins known as effector proteins, which establish a network of interactions critical for their infection process. Our goal is to unravel this network of protein interactions to gain a better understanding of how infections occur and how the immune system responds to different infection strategies. To achieve this, we have created mutant strains of these pathogenic bacteria. We then use these mutants to infect mice and observe the effects they have on the intestine. This allows us to gain insights into the mechanisms behind infection and immunity.
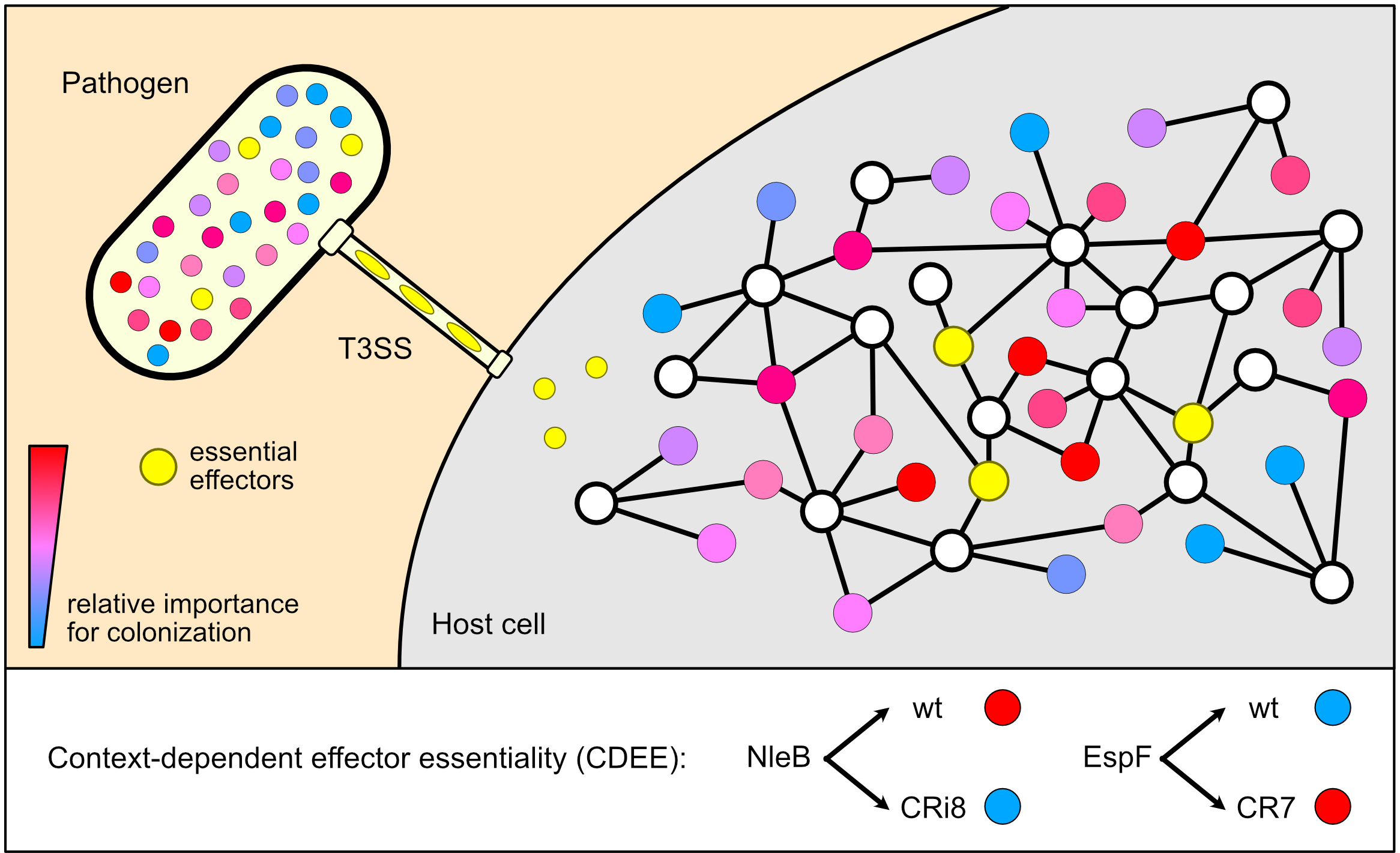
Figure 1. Type III secretion system (T3SS) effectors form interconnected networks in vivo. Pathogens inject T3SS effector proteins into host cells in order to subvert cellular processes and establish a successful infection. Once infected, these effectors (depicted as colored circles) can form connections (edges) with host proteins (white circles) and other effectors, building a robust network that can withstand significant perturbations without affecting virulence. One host protein can connect to more than one effector by edges with positive/stimulatory and/or negative inhibitory nature; for example, T3SS effectors NIeB and EspL in attaching and effacing (A/E) pathogens can inhibit RIPK1 by modifying its death domain or cleaving its RHIM domain respectively, while the Salmonella guanine nucleotide exchange factors (GEFs) SopE and SopE2 can activate Rac1 and Cdc42, while SptP, a GTPase-activating protein (GAP), can inactivate these GTPases. Aside from effectors that are essential in all networks (shown in yellow), the importance of each effector for in vivo colonization depends on the connections it establishes within the subnetwork; here, the color of each effector indicates its relative importance for colonization. This phenomenon is termed context-dependent effector essentiality (CDEE), here exemplified by NleB, which is required for colonization in the wild-type (WT) subnetwork (in red) but becomes dispensable in CRi8 (in blue), and EspF, which becomes necessary for colonization in the context of the CR7 subnetwork (in red).
Furthermore, we aim to harness the power of probiotics for biotechnological applications in the field of biomedicine. Using advanced techniques from synthetic biology, we intend to develop methods for manipulating strains of bacteria that were previously not well-understood. These particular bacterial strains have the potential to serve as indicators of good health and offer protection against intestinal diseases. Our research seeks to boost the capabilities of these probiotics as valuable tools for improving human health.
| Last Name | Name | Laboratory | Ext.* | Professional category | |
|---|---|---|---|---|---|
| Ruano Gallego | David | 104 | 4678 | druano[at]cbm.csic.es | Investigador |
| Bueno Alonso | Marina | 104 | 4678 | marina.buenoa@estudiante.uam.es | Estudiante TFG |




















